Key Takeaways
- Education ranks among the top 10 fields in terms of data storage needed, with institutions dedicated to research in genomics and other physical sciences experiencing a particularly acute problem with storing massive amounts of data.
- Five years after it opened, Columbia University's Center for Computational Biology and Bioinformatics (C2B2) found itself facing a data-storage tipping point: emerging high-throughput biological technologies required new storage solutions that could also streamline researchers' access to data.
- To manage its data explosion, C2B2 dramatically redesigned its storage system to independently scale performance and capacity by leveraging "scale-out" storage platforms.
Much talk today focuses on "big data," and for good reason. Big data challenges an IT department's abilities to capture and store data and provide users with reliable, fast access to information without spreading IT resources too thin. Recent research illustrates the growing big data challenge:
- With a collective 269 petabytes of data, education was among the U.S. economy's top 10 sectors storing the largest amount of data in 2009, according to a McKinsey Global Institute survey.
- The world will generate 1.8 zettabytes of data this year alone, according to IDC's 2011 Digital Universe survey.
- Worldwide data volume is growing a minimum of 59 percent annually, a Gartner report estimates.
 Author John Wofford talks about Columbia University's C2B2 (2:26 minutes):
Author John Wofford talks about Columbia University's C2B2 (2:26 minutes):
The situation is particularly acute in university and commercial centers dedicated to genomics and other physical sciences. Every day at Columbia University's Center for Computational Biology and Bioinformatics (C2B2), our computational systems process terabytes of biomedical data. Whether it is to analyze the deluge of next-generation sequence data generated by modern gene sequencing instruments in search of genetic clues for cancer research; to use medical databases to track the spread of disease; or to better understand proteins through biophysical simulation, our computational research all has one thing in common — an ever-growing demand for data availability and capacity. Our researchers' work can lead to new drug discoveries and advances in basic science. But to facilitate this research, we need effective ways to scale data storage capacity and performance.
Since its founding in 2003, C2B2 has evolved from a disparate set of small research labs with separate, variously sized computing and storage systems into a unified research center for computational biology and bioinformatics. As C2B2 grew, so too did our computing and storage systems.
We were forced early on to face some major challenges in storage design, and in the end we have had remarkable success. We have successfully scaled multiple orders of magnitude in both capacity and performance to facilitate our research. Over the past couple of years, I have spoken with several institutions about storage design — several of whom have emulated our architecture — and I want to share with you some of the considerations that led to our current design, as well as some key takeaways for designing scalable, high-performance storage systems today.
C2B2 Storage Challenges
By late 2007, we faced a tipping point in terms of computational and storage performance and capacity. The various high-throughput biological technologies — especially gene sequencers — were becoming more readily available, and we faced a deluge of incoming data that needed to be stored and analyzed. High-throughput biological technologies generally refer to biomedical research "wet lab" equipment that interfaces with computing equipment to rapidly generate research data. Gene sequencers, for instance, take high-resolution images of specially treated genetic material, which can be used to deduce the sequence of nucleotides in a sample of DNA or RNA. A modern gene sequencer can generate multiple terabytes of image files a week. All this data needs to be analyzed, and researchers processing this data with various bioinformatic algorithms must try to extract scientifically meaningful information.
Our largest computing system held about 400 CPU cores backed by roughly 30 terabytes (TB) of data storage provided by traditional network attached storage (NAS) systems and simple file servers. It was becoming painfully obvious that this system was not just insufficient, it was too small by an order of magnitude. We set our sights on growing our infrastructure into a high-performance, Linux-based computer cluster of roughly 4,000 CPUs, which would offer more than 10 times our existing computing performance.
We also knew we needed on the order of 200 to 300 TB of disk capacity immediately, and we needed enough disk performance to make that data regularly available to those 4,000 CPUs. More importantly, we saw no end to data growth. We needed to design a storage system that would not only meet our needs in 2007 but also scale to meet our needs in the future.
The Options: Traditional NAS, Do-It-Yourself, or Scale-Out NAS?
When considering storage alternatives, we needed to meet our performance and scalability demands while also ensuring reliability and cost effectiveness. We operated with five basic requirements:
- Performance: The system needed to handle the heavy concurrent I/O from a large-scale compute cluster.
- Scalability: With exponential data growth expected, we needed to be able to scale both capacity and performance effortlessly.
- Reliability: System failures can cost researchers time and money, and ultimately they endanger potential breakthroughs. We needed a system with redundancy and reliability built in.
- Manageability: We are a small IT team, and we needed to keep it that way. We required a system we could deploy and maintain without adding a team of storage engineers.
- Cost-effective: We operate primarily on limited, grant-based funds. We needed a system that would be cost-effective both at purchase time and going forward.
Table 1 shows the storage system options we considered, along with their pros and cons.
Table 1. Storage Options: Pros and Cons
| Storage Option | Example | Pros | Cons |
|---|---|---|---|
| Traditional, non-clustered NAS systems | NetApp NAS filers | Well-established technology with products from multiple vendors | Architecture couldn't effectively support high level of concurrent I/O |
| Build our own computing file system | Lustre | Custom-built to meet our specific compute/storage needs; supports high level of concurrent I/O | Management and lack of vendor support would strain IT department's resources; no assurances of reliability |
| Scale-out NAS | EMC Isilon | Easy to scale in performance and capacity; handles high level of concurrent I/O; easy to manage; reliability and service backed by vendor support | At the time, clustered NAS was still a relatively new technology
|
Traditional NAS Systems
We began our decision-making process by considering systems like the traditional NAS systems we already maintained. Traditional NAS systems typically consist of one or more NAS head controllers, which manage a file system stored on a storage area network (SAN). A traditional NAS is a natural outgrowth of a simple file server. One piece of hardware manages all of the storage requests, but you distribute the storage over many disks in the back end.
We were already having some difficulty scaling performance of our existing NAS systems, but if you have a technology in place that's working well — but not well enough — it makes sense to at least question whether it can be improved or expanded somehow to meet your needs. We had several conversations about growing the traditional NAS system we were already using. There were four things in the technology's favor:
- NAS systems scale storage capacity with relative ease by adding new back-end SAN nodes.
- It's a solution we knew and had experience with.
- NAS systems are a time-tested data center staple: something everyone in the enterprise IT field is familiar with.
- Because of their wide popularity, NAS systems were available from a variety of major vendors in a variety of flavors.
To a degree, we knew from experience that continuing with traditional NAS would be a safe bet. After all, no one ever gets fired for buying the industry standard. We could have purchased a giant traditional NAS system and it would have worked — it just wouldn't have worked as well or as cost-effectively as other alternatives. Capacity scaling isn't a challenge per se, but performance scaling can be problematic.
When you look at the architectures of high-performance computing and traditional NAS from a performance standpoint, there is a fundamental design disconnect. High-performance computing is all about massive parallel processing, while traditional NAS systems are designed to filter everything through a single NAS head controller. We knew it wouldn't be unusual for us to have 4,000 tasks asking for storage access at the same time. Passing all that storage I/O through a central NAS head controller is not a sensible approach. The architecture isn't designed for that level of I/O demand, and scaling for higher demand would potentially require costly replacement of the NAS head. We could postpone the scalability issue temporarily by engineering a very large system from the beginning, but that would not be cost-effective. We decided that traditional NAS was the wrong design for our needs.
The DIY Approach
Instead of traditional NAS, we focused on a second option: taking the "do it yourself" (DIY) approach to build our own clustered file system using Lustre (a parallel distributed file system) or Global File System (a shared-disk file system for Linux compute clusters), and other similar options. Clustered file systems were designed with demanding parallel workloads in mind and have been a mainstay in high-performance computing for years. A properly designed clustered file system can also offer very high performance. In fact, most of the biggest supercomputers out there are backed by some form of clustered file system.
Additionally, I'd had experience in this area previously. I had worked on a project at Los Alamos National Laboratory (LANL) where we had experimented with various clustered and scalable file systems. As part of that project, I had developed an InfiniBand-based network file system that achieved impressive storage performance for minimal cost. (This project won the Supercomputing Storage Challenge, small systems class, at the Supercomputing 2007 conference.1) Because the DIY approach uses commodity hardware and, typically, cheap or free software, these systems can often be built for considerably less than other commercial options.
We experimented with a clustered file system designed around our requirements and considered how this option stacked up to our five requirements. Though pleased with the performance, purchase cost, and scalabilty, we realized the DIY approach wouldn't be feasible financially in the long run due to the increased management and support overhead.
Clustered file systems are designed with performance in the forefront. In many installations where they are used, these systems are treated exclusively as computing "scratch space" — i.e. high-performance space with little focus on reliability. We needed more from our storage solution's reliability. But we couldn't afford to have team members spending their time examining storage packet traces and performing low-level analysis of storage system problems whenever they occurred. We determined we would need to add one to two storage engineers to deliver the performance and reliability we required, which drove the total cost of ownership too high.
Scale-Out NAS
Finally, we looked at what was a relatively new technology on the market at that time: scale-out, or clustered, NAS. With scale-out NAS, you have a distributed file system that runs concurrently across multiple, clustered NAS nodes. It provides access to all files from any of the nodes, regardless of the file's physical location. The architecture is similar in many ways to the DIY clustered file system, with the added benefit of enterprise features and a design focus on more than performance.
Scale-out NAS was a rather risky investment at that time, however. Unlike the other two options, scale-out NAS was a fairly new technology in 2007, and its limitations and capabilities weren't well known. Very few vendors offered scale-out NAS, and they hadn't been around long, so there was a chance we'd be buying from a company that might fold. Also, it was difficult to ascertain which vendors had the best implementation. There wasn't much in the way of guarantees, which gave us pause. Not surprisingly, some faculty at C2B2 had concerns that we would be investing deeply in a relatively untested technology from an unknown company.
As it happens, the storage system I had designed at LANL shared some remarkable similarities with some of the scale-out NAS systems, which put me in a unique position to assess the technologies. After grilling the vendors and a couple of existing customers in some technical detail on the storage subsystem, I was convinced that the technology was well designed and here to stay.
Ultimately, we decided that scale-out NAS would give us all the benefits of the DIY clustered file system with the added reliability, management, and support we were looking for, which made this option the best fit for our criteria. Our decision proved to be the right one. Nearly five years later, a wealth of technical data, market data, and use cases support the validity of scale-out NAS. Today, almost every major storage vendor on the market offers some form of scale-out NAS solution, and these systems are being successfully deployed for almost every storage application. Figure 1 shows a diagram of our multi-tiered data storage architecture.
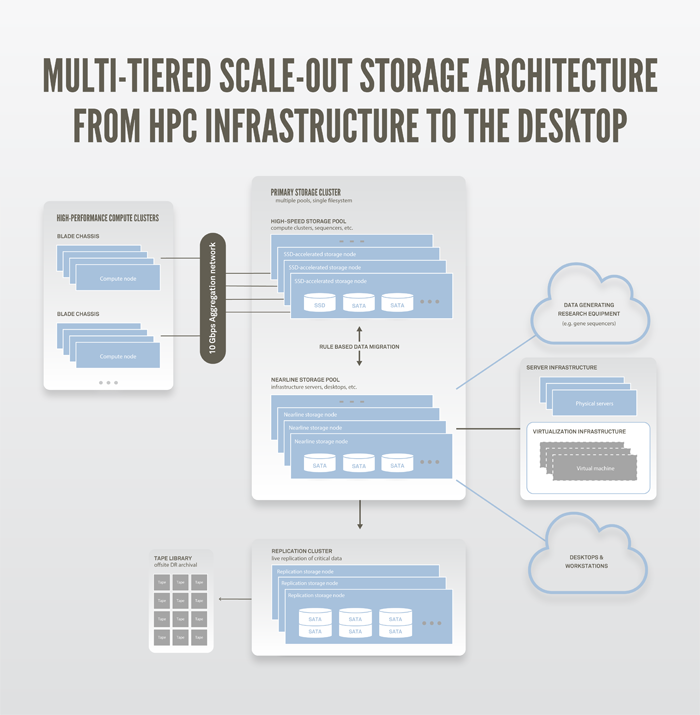
Figure 1. C2B2 storage architecture
Design Considerations for Scale-Out NAS
Given five years of deployment of our scale-out NAS, and having worked with several other institutions on their storage design, we have learned a few things about what makes these deployments successful. In the following sections I will go over three specific problems we faced and how we used modern storage systems to plan for and solve them. Although these challenges are related to high-performance computing, the basic takeaways are the same for many environments.
1. Build Small
On any given day, we have 120 or more users running thousands of computational jobs, from molecular biophysical simulations to sequence analysis and genome assembly, against our storage system. Since we deal with many different users performing a variety of tasks, it's impossible to predict exactly what the total storage workload will be. Without a well-defined workload, it's extremely difficult to predict which system specifications are appropriate for the environment. In place of systems engineered for specific environments, we must design on basic heuristics for guessing a storage design, and in my experience many users find themselves in the same situation.
Let's look at one such heuristic for high-performance computing workloads. Imagine that you're trying to pair storage with a 500-node compute cluster (i.e., 500 computational servers). If you want network storage access to approximate the storage performance of a local disk, you might say that you need one disk for each node. This is a common approach and in general will achieve good results. But it's also, in most cases, overkill.
Part of the beauty of scale-out architecture is the ability to scale performance and capacity as easily as plugging in a new piece of hardware. This allows a different kind of approach to sizing a system. When we designed our initial deployment of scale-out hardware, we intentionally chose a configuration that was a little less than what we needed. This allowed us to get a real system in place, throw real workloads at it, and add hardware until we were meeting our demands.
As expected, the initial configuration couldn't quite stand up to our full computational power. A few months later, we added more scale-out storage nodes and were able to painlessly scale up to meet demands. Incidentally, we were able to make do with far fewer disks than we might have initially guessed, and that ultimately saved us money. The bottom line: In the scale-out world, you can buy a minimal configuration and add performance and capacity separately, as needed.
2. Build Once
From a technical standpoint, the scale-out NAS deployment was one of the easiest implementations I've ever done. We spent more time discussing how to organize all of our data in one directory structure than we did on how to make the scale-out NAS system actually work. All told, it took about a month and a half to transfer all of our data from the traditional NAS system to the new scale-out NAS system. On the other hand, the hardware deployment was simple. Our initial install went from the pallet to serving data in one afternoon. Subsequent node additions have been even more effortless. Even removing old hardware proves painless. Today, we run a completely refreshed set of hardware on the same file system we ran in 2008. In figure 2 the solid orange line indicates the approximate storage requirement, while the dashed red line indicates growth of the physical storage system.
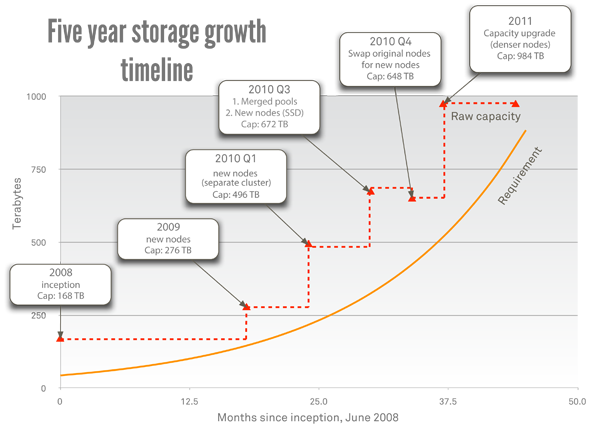
Figure 2. Timeline of C2B2's five-year storage growth
In the old world of storage design, when we deployed a new system we always needed a similar data planning and migration phase. Fortunately, scalable storage changes that model. We haven't had to migrate data from one file system to another since that initial migration, and that's a good thing. As data sets grow, this kind of data migration is untenable. For instance, our data went from 30 TB to nearly 1 PB in the past few years (see figure 2). A migration and reorganization of data today would be a substantial undertaking. Scalable storage gives the option to set up one storage system and scale it indefinitely. The key is to design an organized, scalable structure for your data from the beginning and scale that system up rather than replacing it.
3. Take Advantage of Diverse Hardware
By 2010, we thought we had more or less hit on a storage design that was going to give us all of the performance we needed. Unfortunately, despite having adequate storage processing, cache, network throughput and disk, we started getting very slow response time from our system. The slow response did not seem to be directly correlated to either computational or storage system load.
Fortunately, we were able to use some vendor-supplied analytics software to take a look at the workload of our system. We discovered that more than 40 percent of our total workload consisted of namespace read operations (figure 3 shows protocol operations per second over one week; the heatmap below shows the distribution of protocol operations by operation class). Namespace reads are operations that, rather than accessing data, query information about the file system. This happens, for instance, when you list the contents of a directory. Further investigation revealed that this was being caused primarily by certain bioinformatics programs that query "databases" of tens of thousands of files, which are "indexed" by their filename. We had thousands of these running at once, and while our storage was handling reads and writes quite well, it couldn't handle the namespace operations at this rate.
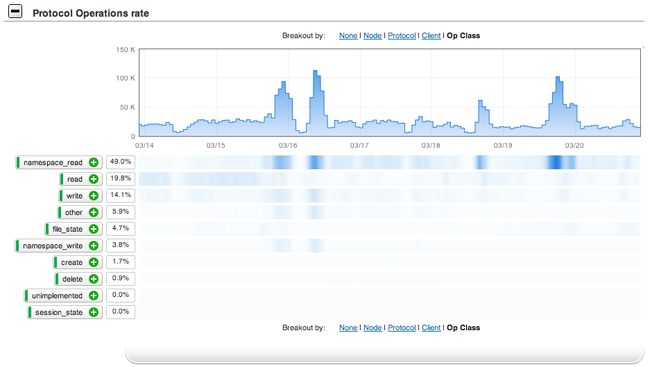
Figure 3. Heatmap of file system operations
Since traditional disks store data on spinning platters, there is a typical delay time — known as seek time — to access nonsequential data. Namespace operations require a lot of nonsequential disk reads, which means that they are constrained by seek time. Solid-state disks (SSDs), which are now quite prevalent, don't use spinning platters, so their seek time is only limited by how fast they can transfer data to memory buffers. Various benchmarks report that SSDs can perform seeks as much as 40 times faster (or more) than the fastest magnetic disks. We worked with our vendor to deploy storage nodes that would include a small number of SSDs on each node to store the file system namespace information specifically (see figure 4). The end result was an immediate, dramatic improvement in file system responsiveness.
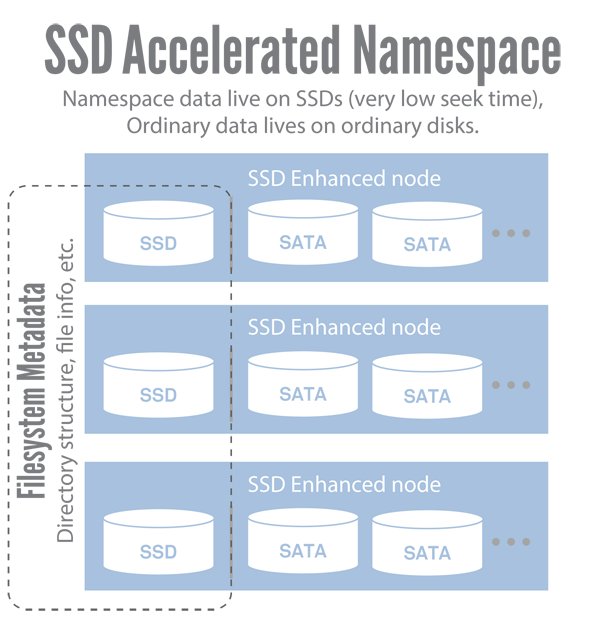
Figure 4. SSD accelerated namespace
Although namespace reads may not be a problem for everyone, the basic point is that different workloads are best handled by different hardware, and there is a lot of hardware diversity out there. While many traditional solutions are optimized to handle only homogeneous hardware, some scale-out storage options allow the flexibility to install new nodes that can meet new storage challenges as they happen. In fact, with the multi-tiered storage options that some vendors offer these days, one can even scale different kinds of hardware independently. In these systems, one can have multiple kinds of storage node in the same file system. These combined capabilities allow modern systems to adapt to use modern hardware as needed, whether SSDs, the latest high-capacity disk, or the latest network technology.
Today and Going Forward
To date, we have installed roughly 1 PB of storage in 18 nodes of three different types. This design has provided us cost-effective and manageable storage capacity and performance that can scale with us as we grow. Today we find ourselves at a tipping point much like we were five years ago. We hope to scale our computing performance by another order of magnitude and more than double our storage capacity over the next year. This time, however, we do not foresee major architectural changes to our storage environment. We expect to deploy new storage to meet our increased computing and capacity demands without data migration or major infrastructure changes. We will just scale up where we need it.
- Michael S. Warren and John Wofford, "Astronomical Data Analysis with Commodity Components," recipient of the Best Research in High Performance Computing, Small System Storage Challenge category, at the SC07 conference.
© 2012 John Lowell Wofford. The text of this EDUCAUSE Review Online article (September/October 2012) is licensed under the Creative Commons Attribution-Noncommercial-ShareAlike 3.0 license.
